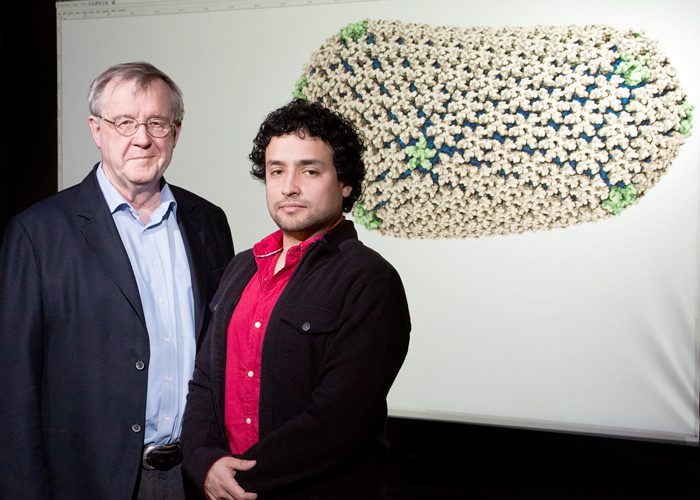
Using GPU-based simulations researchers at the Beckman Institute, University of Illinois have solved the entire molecular structure of the HIV capsid (protein shell). X-ray crystallography is used to probe the structure of the proteins that make up a virus. This is however inadequate when trying to determine how those proteins are assembled to build the virus. The solution: simulate the process with data from both ends of the process using high powered computers. This is exactly what the researchers did using the Blue Waters supercomputer at the University of Illinois. The machine has 237 Cray XE6 cabinets and 32 Cray XK7 cabinets with Nvidia Tesla Kepler GPU cluster. This provided simulations for detailed molecular motion on the 1300 identical proteins contained in the capsid.
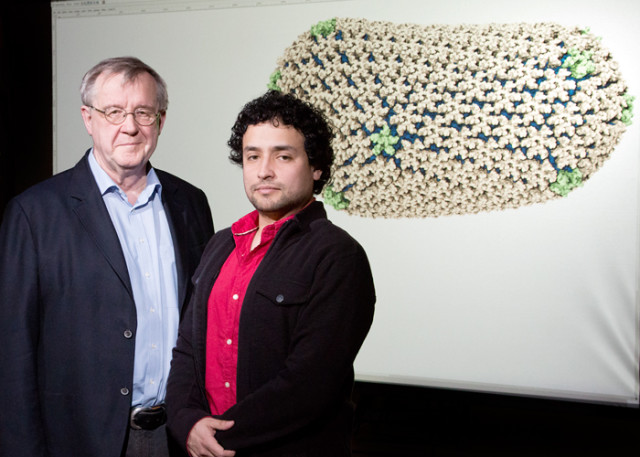

Although capsid proteins tend to form hexamers and pentamers, the HIV virus has an asymmetrical form. Many viruses also do have a variety of stable structures they can assume. From a simulation of 64 million atoms, the experiment determined that the capsid contains 216 hexamers and 12 pentamers. When moving between cell hosts, the capsid protects the virus. Inside a cell, it flexes open in order to attack the genetic matter of the cell host. There are currently no anti-capsid drugs developed for the HIV virus where the same exist for other viruses. Understanding how the HIV capsid is assembled makes it easier to develop new drugs which cause premature opening of the capsid, or perhaps block its opening altogether.
The study utilized NAMD, an open-source dynamics package on the GPU cluster. NAMD is written in an efficient parallel programming language known as Charm++. The study has been published under the title “Structure of the Mature HIV-1 Capsid by Cryo-EM and All-Atom Molecular Dynamics Simulation”.